
Biology
Whether you’re preparing for a career in healthcare, conservation, or biotech, biology students benefit from experiences in our state-of-the-art labs in the Billie Tisch Center for Integrated Sciences, the 500+ acre North Woods, and world-class STEM facilities.
What will you learn?
Our curriculum challenges you to understand life at every scale. Foundational coursework explores fundamental molecular and cellular processes, structure-function relationships, behavior and ecology, and evolutionary patterns. Majors pursue specialized concentrations while benefiting from a dynamic liberal arts education. Many also follow a pre-health track.

Four concentrations

Health careers

Advanced research
What will you accomplish here?
At Skidmore, students present at conferences, publish research, and study ecosystems around the world. You'll graduate with the kind of experiences that set you apart. Here are a just a few examples of faculty-student co-authored research publications and presentations, as well as approved biology study abroad programs:
| Conferences | Publications | Study Abroad |
|---|---|---|
|
|
|
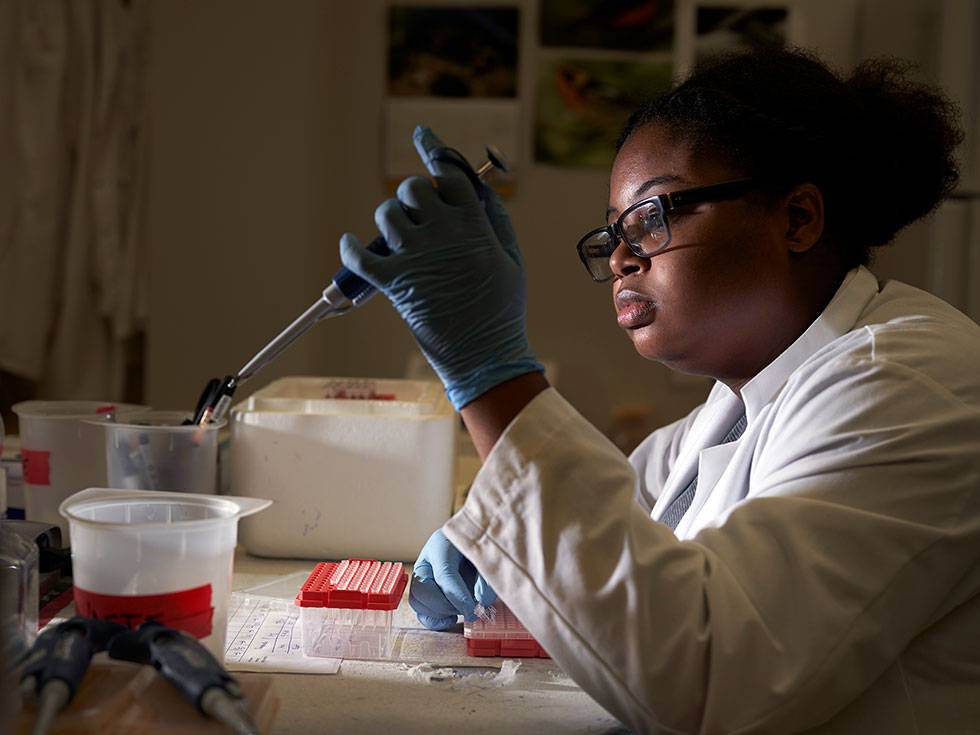


Real-world experience
Where will you go?
Whether you're headed to a hospital, a research lab, or a field station, your Skidmore experience sets you up for real impact and real opportunity.
| Career Paths | Employers | Graduate and Medical Schools |
|---|---|---|
|
|
|
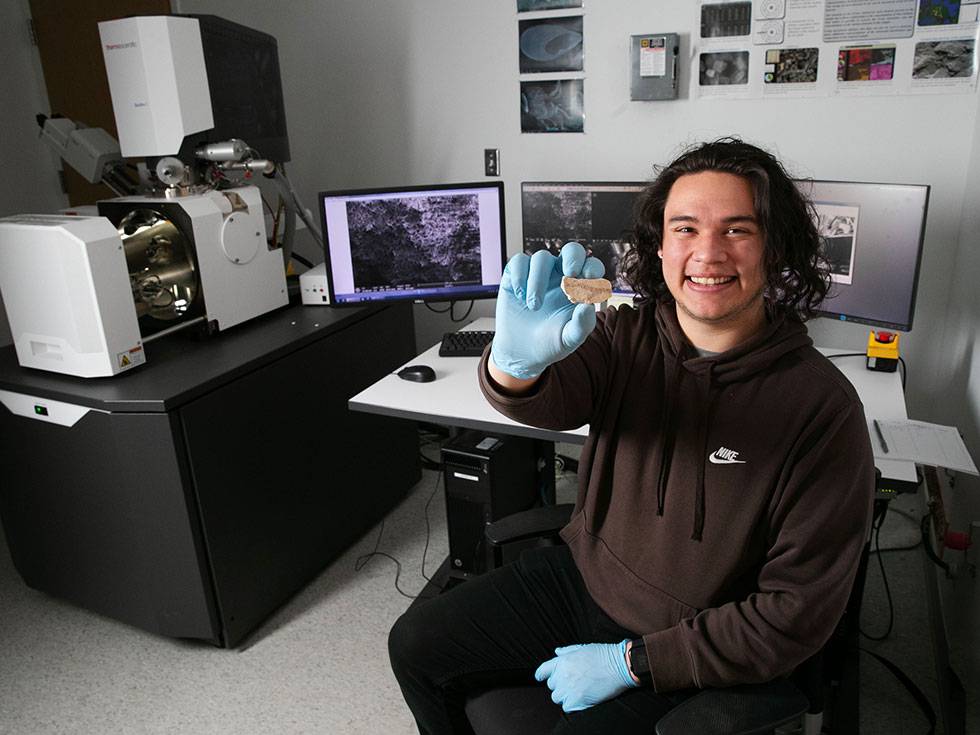

See the world like never before
Skidmore’s McGraw Microscopy Imaging Center gives students access to an impressive collection of microscopes rivaling major research institutions.
Beyond the lab
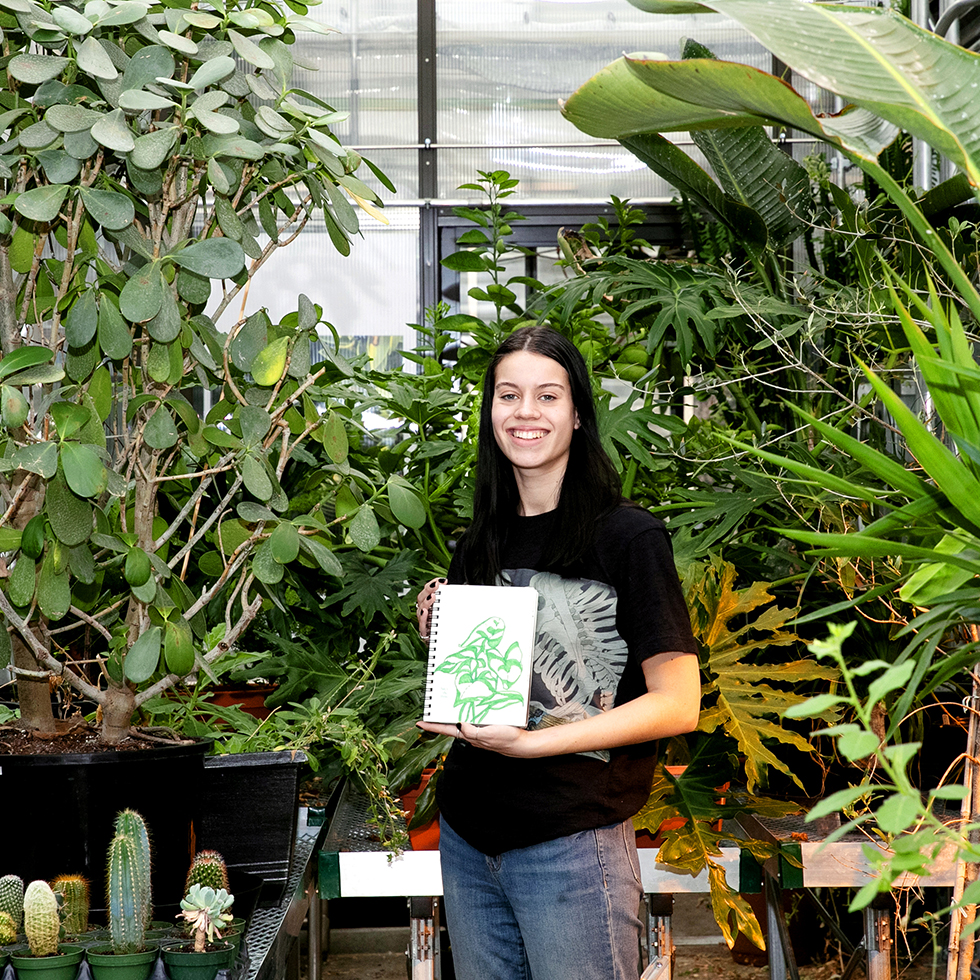

The art of science
Why not both?
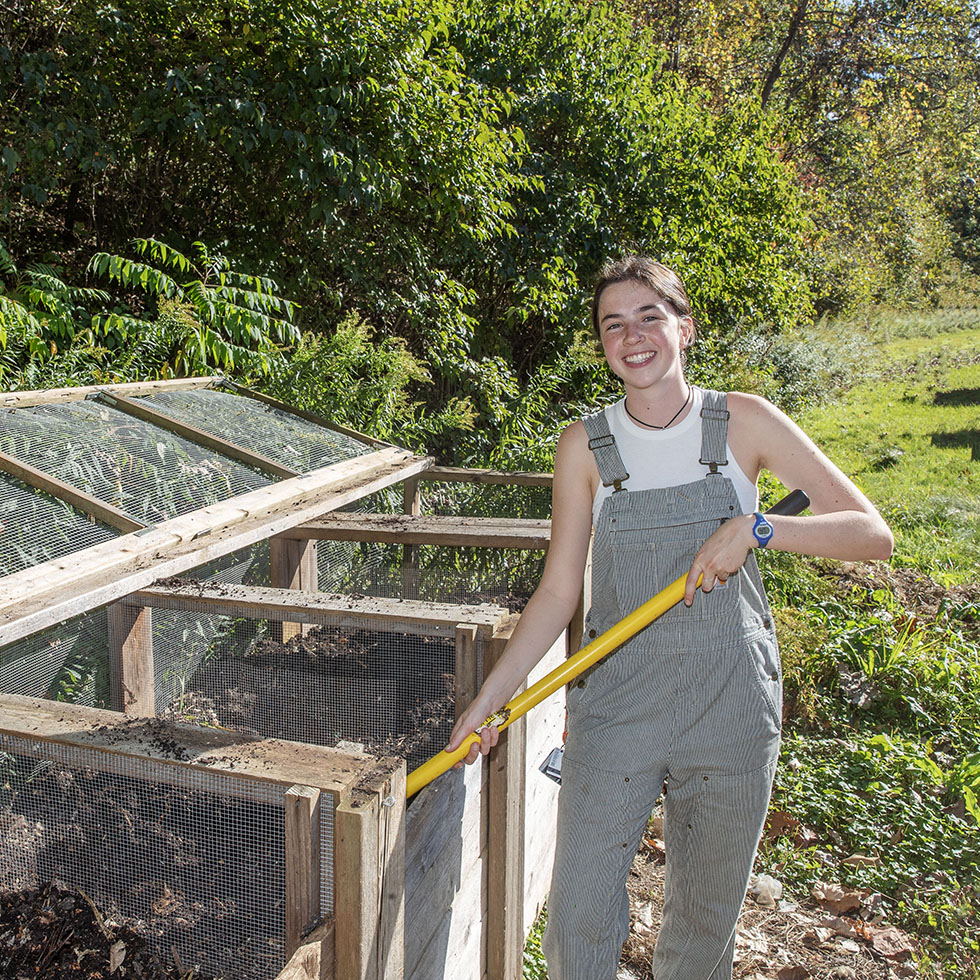

Biology meets sustainability
